Scientists at Sanford Burnham Prebys demonstrated that vesicles traveling between cells in the brain carry more complete instructions for altering cellular function than previously thought.
In findings published in Cell Reports, senior author Jerold Chun, MD, PhD, and team also discovered that the biological instructions within these vesicles differed significantly in postmortem brain samples donated from patients suffering from Alzheimer’s disease.
Researchers call the tiny brain bubbles under scrutiny in this study small extracellular vesicles (sEVs). These tiny biological water balloons are produced by most cells in the body to ferry a wide variety of proteins, lipids and byproducts of cellular metabolism, as well as RNA nucleic acid codes used by recipient cells to construct new proteins.
Because this biologically active cargo can easily elicit changes in other cells, scientists are interested in brain sEVs as a medium for passing along normal as well as bungled instructions for misfolded proteins that accumulate in the brain as neurodegenerative diseases such as Alzheimer’s disease progress.
To be a potential contributor to the buildup of unwanted proteins, sEVs would have to carry blueprints with sufficient information to enable other cells to produce the problematic proteins. Most previous research had indicated that the messenger RNA (mRNA) carrying plans for proteins were chopped into too many shorter fragments to allow recipient cells to change their construction patterns.
“We found quite the opposite to be true in our study,” says Chun, professor in the Center for Genetic Disorders and Aging Research at Sanford Burnham Prebys. “We identified more than 10,000 full-length mRNAs through the use of a relatively newer DNA sequencing technique called PacBio long-read sequencing.”

Immunofluorescence was used to validate primary cell culture purity. Image courtesy of the Chun lab.
The team isolated sEVs from the prefrontal cortex of 12 postmortem brain samples donated from patients diagnosed with Alzheimer’s disease and 12 from donors without Alzheimer’s disease (or any other known neurological disease). Nearly 80% of identified mRNAs were full-length, allowing them to be transcribed by recipient cells into viable proteins.
“To corroborate the results of long-read sequencing in the human samples, we also looked at vesicles isolated from mouse cells,” says first author Linnea Ransom, PhD, postdoctoral fellow at Sanford Burnham Prebys. “We found similar averages of between 78% and 86% full-length transcripts in three brain cell types: astrocytes, microglia and neurons.”
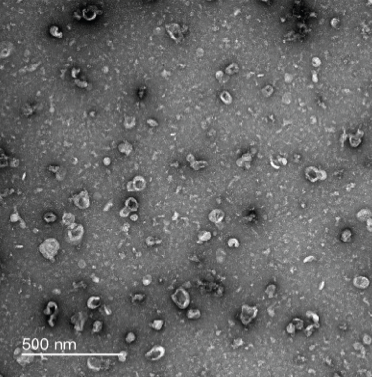
Negative-stain transmission electron microscopy was used to confirm the isolation of small extracellular vesicles in mouse neurons (shown above), as well as in mouse astrocytes and microglia. Image courtesy of the Chun lab.
In addition to analyzing and validating the results regarding the length of mRNAs in brain sEVs, the researchers compared the sequence of genes reflected in the sEV mRNA transcriptome. In Alzheimer’s disease samples, 700 genes showed increased expression whereas nearly 1500 were found to have reduced activity.
The scientists determined that the 700 upregulated genes are associated with inflammation and immune system activation, which fits within known patterns of brain inflammation present in neurodegenerative diseases such as Alzheimer’s disease. The researchers also found many genes associated with Alzheimer’s disease in prior genome-wide association studies also were present in Alzheimer’s disease sEVs.
“The changes in gene expression contained in these vesicles reveal an inflammatory signature that may serve as a window into disease processes occurring in the brain as Alzheimer’s disease progresses,” says Chun.
Following this study, Chun and team will dig deeper into how cells package sEVs and how the enclosed mRNA codes lead to functional changes in other brain cells affected in Alzheimer’s disease. Better understanding of sEVs and their mRNA contents may enable the discovery of biomarkers that could be used to improve early detection of Alzheimer’s disease and potentially other neurological conditions, while identifying new disease mechanisms to provide new therapeutic targets.
“Additionally, sEVs naturally occur as a vehicle for transporting biologically active cargo between cells, so it also may be possible to leverage them as a targeted delivery system for future brain therapies,” says Chun.
Additional authors on the study include Linnea S. Ransom, Christine S. Liu, Emily Dunsmore, Carter R. Palmer, Juliet Nicodemus, Derya Ziomek and Nyssa Williams, all at Sanford Burnham Prebys.
The study was supported by the National Institute on Aging (R01AG065541 and R01AG071465), National Institute of General Medical Sciences (T32GM007752) and Rotary International.
The study’s DOI is 10.1016/j.celrep.2024.114061.