Related Disease
Cancer
Phenomena or Processes
Aging, Cancer Biology
Techniques and Technologies
Bioinformatics, Computational Biology, Drug Discovery, Machine Learning
Developing cancer preventive therapies using the power of AI.
At the heart of our research is the mission to develop cancer-preventive therapies using the power of artificial intelligence. Our current focus lies in dissecting the role of aging in cancer susceptibility, a crucial factor often overlooked in conventional research. We employ computational tools to analyze multi-omics data to understand the aging-induced alterations in the tissue microenvironment that increase cancer risk. But we don’t stop at understanding these changes. We apply this knowledge in designing preventive therapy candidates that specifically target these alterations. Drawing on my experience in machine learning, drug discovery, and precision oncology, our lab is on a quest to reimagine the drug discovery pipeline.
“As we aim to push the boundaries of cancer prevention research, we are hiring individuals eager to contribute to this mission at multiple levels including postdoctoral researchers, experienced computational biologists, and, PhD students.”
Sanju Sinha’s Research Report
My key contributions to the field of computational oncology during my PhD and postdoctoral tenure are outlined below:
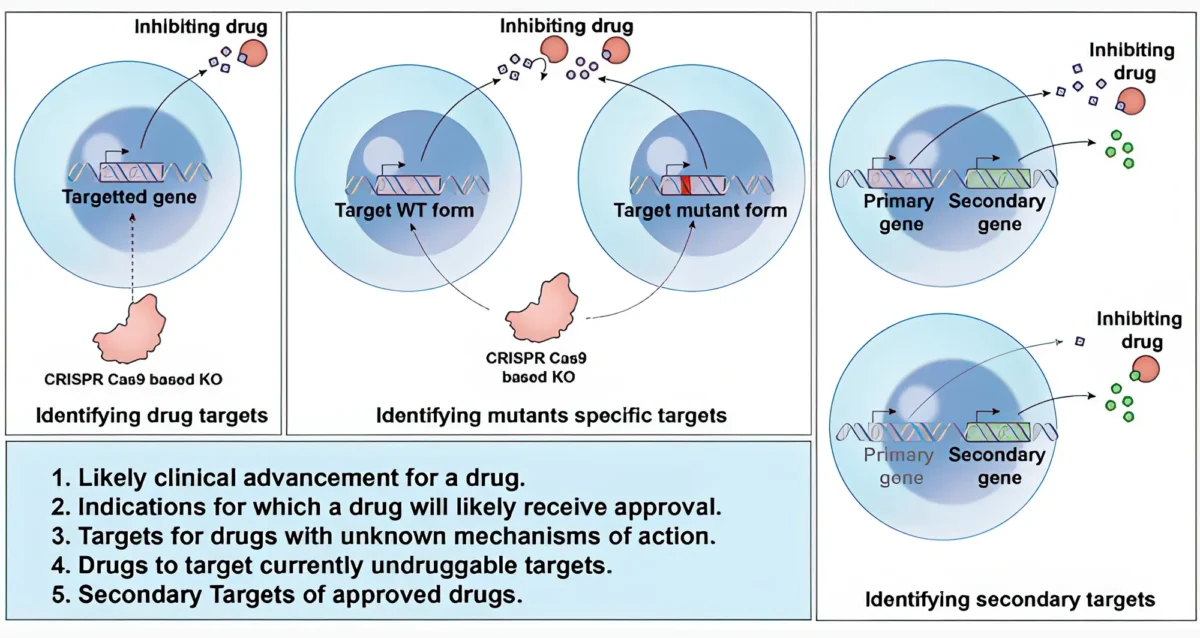
DeepTarget: A Computational Tool for Decoding the Mechanism of Action for Cancer Drugs
DeepTarget, our innovative computational tool, integrates data from genetic and drug screens to intricately understand the mechanism of action of cancer drugs. It identifies both primary and secondary drug targets, paving the way for a comprehensive understanding of drug function, its optimal indications, and clinical potential.
Revealing the Potential Cancer Risks associated with Genetic Editing
Through computational analyses of vast genetic screens, we shed light on the inherent selection potential of specific cancer gene mutations associated with CRISPR-Knockout editing. This work has significantly enhanced our understanding of the risks associated with gene editing.
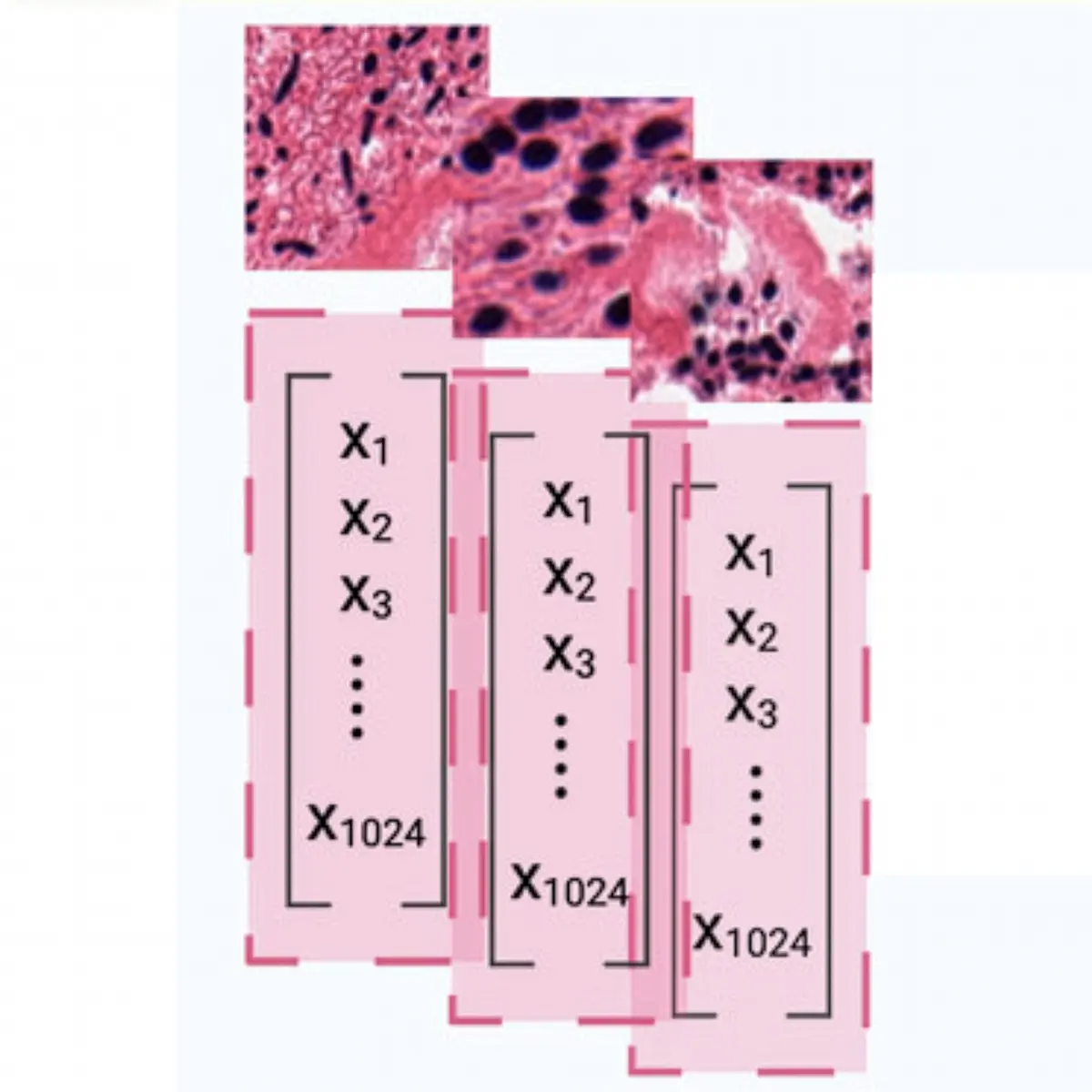
First Single-cell based Precision Oncology Framework: A Proof-of-concept
Our innovative approach to precision oncology utilizes single-cell transcriptomics to predict patient treatment responses and detect resistance. We demonstrated its efficacy using patient-derived primary cells and three recent single-cell clinical cohorts, providing a powerful tool for the next generation of precision oncology approaches.
Why High Tumor Mutation Burden Biomarker of Immunotherapy is only effective for Certain Cancer Types?
We unveiled the microenvironment context that can determine if the high-tumor mutation burden (TMB) biomarker will be effective in a certain cancer type – High M1 Macrophage levels and Low Resting Dendritic Cells. Based on this, we also predicted the rare tumor types with High-TMB where immunotherapy is most likely to be successful.
Uncovering Therapeutic Prospects for African Americans: A Step Forward in Inclusive Cancer Research
Our work revealed distinct biological differences in tumors across African American and European American patients. Most notably, we identified a higher prevalence of homologous recombination deficiency in African American patients, pointing towards promising, personalized treatment options.