Research Model: Clinical and Transitional Research
Dr. Yu Xin (Will) Wang received his PhD at the University of Ottawa where he identified cellular asymmetry and polarity mechanisms regulating muscle stem cell self-renewal and skeletal muscle regeneration. He then carried out postdoctoral training at Stanford University School of Medicine developing single cell multi-omic approaches to characterize the regenerative process and what goes awry with disease and aging.
“I’ve always had a passion for science and became fascinated with how the body repairs and heals itself when I was introduced to the potential of stem cells in regenerative medicine. I was struck by the ability of a small pool of muscle stem cells that can rebuild and restore the function of muscle. My lab at Sanford Burnham Prebys aims to better understanding the repair process and harness our body’s ability to heal in order to combat chronic diseases and even counteract aging.“
Education and Training
Postdoctoral Fellowship, Stanford University School of Medicine
PhD in Cellular Molecular Medicine, University of Ottawa, Canada
BS in Biomedical Sciences, University of Ottawa, Canada
Prestigious Funding Awards
2020: NINDS K99/R00 Pathway to Independence Award
Honors and Recognition
Governor General’s Gold Medal – Canada
Related Disease
Aging-Related Diseases, Amyotrophic Lateral Sclerosis (Lou Gehrig’s Disease), Arthritis, Cachexia, Inflammatory/Autoimmune Disease, Multiple Sclerosis, Muscular Dystrophy, Myopathy, Neurodegenerative and Neuromuscular Diseases, Sarcopenia/Aging-Related Muscle Atrophy, Spinal Muscular Atrophy
Phenomena or Processes
Adult/Multipotent Stem Cells, Aging, Cell Signaling, Development and Differentiation, Epigenetics, Exercise, Extracellular Matrix, Neurogenesis, Organogenesis, Regenerative Biology, Transcriptional Regulation
Anatomical Systems and Sites
Immune System and Inflammation, Musculoskeletal System, Nervous System
Research Models
Clinical and Transitional Research, Computational Modeling, Human Adult/Somatic Stem Cells, Mouse
Techniques and Technologies
3D Image Analysis, Bioinformatics, Cellular and Molecular Imaging, Gene Knockout (Complete and Conditional), Genomics, High Content Imaging, High-Throughput/Robotic Screening, Live Cell Imaging, Machine Learning, Microscopy and Imaging, Proteomics, Transplantation
The Wang lab is interested in elucidating critical cell-cell interactions that mediate the function of tissue-specific stem cells during regeneration and disease, with a focus on how a coordinated immune response can promote regeneration and how autoimmunity impacts tissue function and hinder repair.
Specifically, the Wang lab aims to identify cellular and molecular crosstalk between muscle, nerve, and immune systems to develop targeted therapies that overcome autoimmune neuromuscular disorders and autoimmune aspects of “inflammaging.”
Yu Xin (Will) Wang’s Research Report
The lab’s research is translationally oriented and utilizes interdisciplinary molecular, genetic, computational (machine learning and neural networks), and bioengineering approaches to view biology and disease from new perspectives. We combine multi-omics sequencing and imaging methods to resolve how different cell types work together after injury to repair tissues and restore function. We use a data-driven approach to identify targetable disease mechanisms and, through collaborations with other researchers and clinicians, develop therapies that promote regeneration. Visit our lab website to learn more.
 Jun 12, 2025
Jun 12, 2025Turning back time on muscle stem cells to prevent frailty from aging
Jun 12, 2025Study from Dr. Will Wang’s lab finds a way to restore stem cells from aged muscle to become young again…
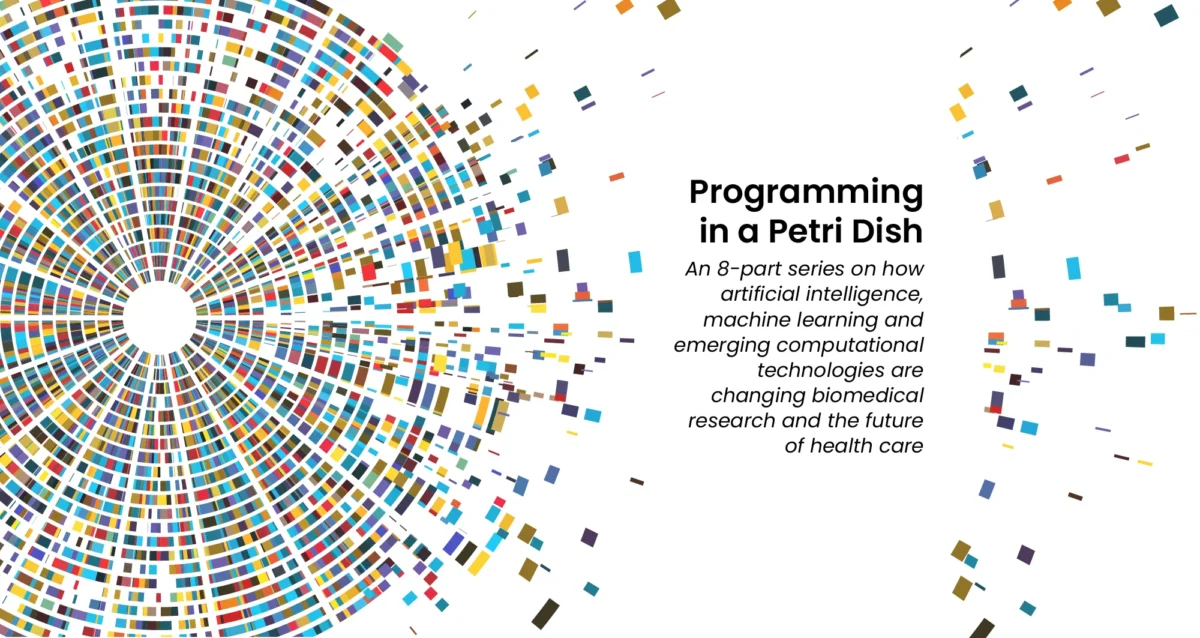 Aug 20, 2024
Aug 20, 2024Mapping the human body to better treat disease
Aug 20, 2024Scientists are investigating the inner workings of our bodies and the cells within them at an unprecedented level of detail.
 Aug 13, 2024
Aug 13, 2024Dodging AI and other computational biology dangers
Aug 13, 2024Sanford Burnham Prebys scientists say that understanding the potential pitfalls of using artificial intelligence and computational biology techniques in biomedical…
 Oct 11, 2023
Oct 11, 2023Inhibiting an enzyme associated with aging could help damaged nerves regrow and restore strength
Oct 11, 2023New research has demonstrated a way to accelerate recovery from peripheral nerve injury by targeting an enzyme that was thought…
 Jan 26, 2023
Jan 26, 2023Three big questions for cutting-edge biologist Will Wang
Jan 26, 2023Will Wang’s spatial omics approach to studying neuromuscular diseases is unique.
 Nov 23, 2022
Nov 23, 2022Yu Xin (Will) Wang joins Sanford Burnham Prebys to advance regenerative medicine
Nov 23, 2022Molecular biologist Yu Xin (Will) Wang, PhD, has joined Sanford Burnham Prebys as an assistant professor in the Development, Aging,…
Pier Lorenzo Puri earned his MD at the University of Rome “la Sapienza” in 1991. Dr. Puri completed his internship in Internal Medicine at the hospital “Policlinico Umberto I” (Rome) from 1992 to 1997, and defended an experimental thesis on the vascular effects of angiotensin II to graduate as Specialist in Internal medicine at the University of Rome “la Sapienza” in 1997. During this time he was frequently working at the Freien University of Berlin, as visiting scientist at the Deprtment of Biochemistry and Molecular Biology, to perform experiments of protein and DNA microinjection in cultured cells. Dr. Puri trained as a post-doctoral fellow at the University of California San Diego (UCSD), in the department of Cell Biology, under the supervision of Dr. Wang, from 1997 to 2001. He was appointed as Staff Scientist at the Salk Institute (La Jolla) in 2001, and became an Assistant Telethon Scientist at the Dulbecco Telethon Institute in Rome in 2002. He was upgraded to Associate Telethon Scientist at the Dulbecco Telethon Institute in Rome since 2007 and became Senior Telethon Scientist, Dulbecco Telethon Institute, in 2012, but declined this position. Dr. Puri joined Sanford Burnham Prebys as an Assistant Professor in 2004. He has been promoted to Associate Professor in 2010 and full Professor in 2015. From 2008 to 2016 Dr. Puri served as Adjunct Professor of Pediatrics at the University of California, San Diego. From 2008 to 2013 Dr Puri was an Associate Member of Sanford Children’s Health Research Center. Dr Puri has been Director of the laboratory of Epigenetics and Regeneration at Fondazione S. Lucia, Roma, Italy, but stepped down this position since 2019.
Education
University of California San Diego, Postdoctoral, Department of Biology
University of Rome La Sapienza, PhD, Internal Medicine
University of Rome La Sapienza, MD, Internal Medicine
University of Rome La Sapienza, Undergraduate, Internal Medicine
Other Appointments
2020-2024: Member of the Science Advisory Board (SAB) European Commission-funded Consortium BIND (Brain Involvement In Dystrophinopathies)
2015-2019: Standing Member, NIH Study Section (SMEP)
2010-present: Member of Editorial Board of Skeletal Muscle
Related Disease
Aging-Related Diseases, Childhood Diseases, Molecular Biology, Muscular Dystrophy, Myopathy, Neurodegenerative and Neuromuscular Diseases, Rhabdomyosarcomas, Sarcopenia/Aging-Related Muscle Atrophy, Spinal Cord Injury, Transcription Factors
Phenomena or Processes
Adult/Multipotent Stem Cells, Aging, Cell Biology, Cell Cycle Progression, Cell Differentiation, Cell Signaling, Cellular Senescence, Development and Differentiation, Disease Therapies, DNA Damage Checkpoint Function, Epigenetics, Gene Regulation, Phosphorylation, Regenerative Biology, Signal Transduction, Transcriptional Regulation
Anatomical Systems and Sites
General Cell Biology, Musculoskeletal System
Research Models
Clinical and Transitional Research, Cultured Cell Lines, Human Adult/Somatic Stem Cells, Mouse Embryonic Stem Cells, Mouse Somatic Stem Cells, Primary Human Cells
Techniques and Technologies
Bioinformatics, Cellular and Molecular Imaging, Gene Expression, Genomics
Puri’s lab group investigates the molecular and epigenetic regulation of gene expression in skeletal muscle progenitors and other muscle-resident cell types (including fibro-adipogenic progenitors, cells from the inflammatory infiltrate, cellular components of neuro-muscular junctions) during physiological and pathological perturbations of skeletal muscle homeostasis.
- We use molecular, biochemical and epigenetic tools to understand structural and functional principles of the 3D genome organization that regulates gene expression during muscle regeneration and diseases.
- A topic of particular interest is the analysis of chromatin interactions that define the 3D genome organization and the identification of structural and functional interactions that regulate cell type-specific patterns of gene expression in response to cues released within the skeletal muscle regenerative environment in health and disease conditions, such as muscular dystrophies and other neuromuscular diseases.
- The knowledge derived from our studies is instrumental to elucidate the pathogenesis of muscular disorders and discover pharmacological interventions that promote muscle regeneration to repair diseased muscles.
- Current translational focus is devoted to:
- the study of the therapeutic potential of HDAC inhibitors for treatment of Duchenne Muscular Dystrophy (DMD)
- the identification of genome variants associated to DMD patient-specific patterns of expression of disease-modifier genes that can account for individual trends of disease progression beyond the common genetic deficiency of dystrophin
- the effect of dystrophin deficiency and restoration by gene therapy on 3D genome and transcriptional output of DMD myofibers; the therapeutic potential of extracellular vesicles released by fibro-adipogenic progenitors of DMD skeletal muscles exposed to HDACi.

Puri Lab
Pier Lorenzo Puri’s Research Report
Puri’s lab group investigates the molecular and epigenetic regulation of gene expression in skeletal muscle progenitors and other muscle-resident cell types (including fibro-adipogenic progenitors, cells from the inflammatory infiltrate, cellular components of neuro-muscular junctions) during physiological and pathological perturbations of skeletal muscle homeostasis.
We use molecular, biochemical and epigenetic tools to understand structural and functional principles of the 3D genome organization that regulates gene expression during muscle regeneration and diseases
A topic of particular interest is the analysis of chromatin interactions that define the 3D genome organization and the identification of structural and functional interactions that regulate cell type-specific patterns of gene expression in response to cues released within the skeletal muscle regenerative environment in health and disease conditions, such as muscular dystrophies and other neuromuscular diseases.
The knowledge derived from our studies is instrumental to elucidate the pathogenesis of muscular disorders and discover pharmacological interventions that promote muscle regeneration to repair diseased muscles
Current translational focus is devoted to:
- the study of the therapeutic potential of HDAC inhibitors for treatment of Duchenne Muscular Dystrophy (DMD)
- the identification of genome variants associated to DMD patient-specific patterns of expression of disease-modifier genes that can account for individual trends of disease progression beyond the common genetic deficiency of dystrophin
- the effect of dystrophin deficiency and restoration by gene therapy on 3D genome and transcriptional output of DMD myofibers; the therapeutic potential of extracellular vesicles released by fibro-adipogenic progenitors of DMD skeletal muscles exposed to HDACi.
1. Epigenetic regulation of skeletal myogenesis by histone acetyltransferases and deacetylases
Our earlier identification and characterization of acetyltransferases p300/CBP and PCAF, as transcriptional co-activators, and the histone deacetylases HDACs, as transcriptional co-repressors, of the myogenic determination factor MyoD1-3, respectively, inspired the experimental rationale toward exploiting pharmacological inhibition of HDAC to promote skeletal myogenesis.
- p300 is required for MyoD-dependent cell cycle arrest and muscle-specific gene transcription. Puri PL, Avantaggiati ML, Balsano C, Sang N, Graessmann A, Giordano A, Levrero M. EMBO J 1997 Jan 15 ;16(2):369-8
- Differential roles of p300 and PCAF acetyltransferases in muscle differentiation. Puri PL, Sartorelli V, Yang XJ, Hamamori Y, Ogryzko VV, Howard BH, Kedes L, Wang JY, Graessmann A, Nakatani Y, Levrero M. Mol Cell 1997 Dec ;1(1):35-45
- Class I histone deacetylases sequentially interact with MyoD and pRb during skeletal myogenesis. Puri PL, Iezzi S, Stiegler P, Chen TT, Schiltz RL, Muscat GE, Giordano A, Kedes L, Wang JY, Sartorelli V. Mol Cell 2001 Oct ;8(4):885-97
- Stage-specific modulation of skeletal myogenesis by inhibitors of nuclear deacetylases. Iezzi S, Cossu G, Nervi C, Sartorelli V, Puri PL. Proc Natl Acad Sci U S A 2002 May 28 ;99(11):7757-62
- Deacetylase inhibitors increase muscle cell size by promoting myoblast recruitment and fusion through induction of follistatin. Iezzi S, Di Padova M, Serra C, Caretti G, Simone C, Maklan E, Minetti G, Zhao P, Hoffman EP, Puri PL, Sartorelli V. Dev Cell 2004 May ;6(5):673-84
2. HDAC inhibitors as pharmacological intervention in DMD and other muscular dystrophies
Puri lab discovered that dystrophin-activated nNOS signalling controls HDAC2 activity, thereby revealing a previously unrecognized link between constitutive activation of HDAC2 and alteration of the epigenetic landscape of dystrophin-deficient muscles6,7. This discovery established the rationale for using HDAC inhibitors to counter the progression of Duchenne muscular dystrophy (DMD), by correcting aberrant HDAC activity in dystrophin-deficient muscles8-11.
- Functional and morphological recovery of dystrophic muscles in mice treated with deacetylase inhibitors. Minetti G. C., Colussi c., Adami R., Serra C., Mozzetta C., Parente V., Illi B., Fortuni S., Straino S., Gallinari P., Steinkhuler C., Capogrossi M., Sartorelli V., Bottinelli R., Gaetano C., Puri P.L. Nat. Med 2006 Oct ;12(10):1147-50.
- A Common Epigenetic Mechanism Underlies Nitric Oxide Donors and Histone Deacetylase Inhibitors Effect in Duchenne Muscular Dystrophy. Colussi C., Mozzetta C., Gurtner A. , Illi B., Straino S., Ragone G., Pescatori M., Zaccagnini G., Rosati G., Minetti G., Martelli F., Ricci E., Piaggio G., Gallinari P., Steinkulher C., Capogrossi M.C., Puri P.L*, Carlo Gaetano. Proc Natl Acad Sci U S A 2008 Dec 9 ;105(49):19183-7 *Corresponding author.
- Fibroadipogenic progenitors mediate the ability of HDAC inhibitors to promote regeneration in dystrophic muscles of young, but not old mdx mice. Mozzetta C., Consalvi S., Saccone V., Tierney M., Diamantini A., Mitchel K.J., Marazzi G., Borsellino G., Battistini L., Sassoon D., Sacco A., Puri P.L. EMBO Mol Med 2013 Apr ;5(4):626-39
- Histological effects of givinostat in boys with Duchenne muscular dystrophy. Neuromuscul Disord. Bettica P, Petrini S, D’Oria V, D’Amico A, Catteruccia M, Pane M, Sivo S, Magri F, Brajkovic S, Messina S, Vita GL, Gatti B, Moggio M, Puri PL, Rocchetti M, De Nicolao G, Vita G, Comi GP, Bertini E, Mercuri E. Neuromuscul Disord 2016 Oct ;26(10):643-649
- HDAC inhibitors tune miRNAs in extracellular vesicles of dystrophic muscle-resident mesenchymal cells. Sandonà M, Consalvi S, Tucciarone L, De Bardi M, Scimeca M, Angelini DF, Buffa V, D’Amico A, Bertini ES, Cazzaniga S, Bettica P, Bouché M, Bongiovanni A, Puri PL, Saccone V. EMBO Rep 2020 Sep 3 ;21(9):e50863
- Determinants of epigenetic resistance to HDAC inhibitors in dystrophic fibro-adipogenic progenitors. Consalvi S, Tucciarone L, Macrì E, De Bardi M, Picozza M, Salvatori I, Renzini A, Valente S, Mai A, Moresi V, Puri PL. EMBO Rep 2022 Jun 7 ;23(6):e54721
3. Control of chromatin structure in muscle cells by regeneration-induced signaling pathways
Upon the discovery and characterization of intracellular signaling pathways (i.e. p38, ERK and AKT cascades) that regulate muscle gene expression in myoblasts, in earlier studies during Puri’s postdoctoral training, Puri lab has revealed the mechanism by which muscle environmental cues are converted into epigenetic changes that regulate gene expression in healthy and diseased muscles, via extracellular signal-activated kinase targeting of chromatin-modifying enzymes. These studies provided the first evidence that regeneration activated p38 and AKT signaling cooperatively direct assembly and activation of histone acetyltransferases and chromatin remodeling SWI/SNF complex at myogenic loci in muscle progenitors12,13,15. Moreover, we discovered that regeneration-activated p38 targets Polycomb Repressory Complex (PCR2) at Pax7 locus to promote formation of repressive chromatin during satellite cells a ctivation14.
- p38 Pathway Targets SWI/SNF Chromatin Remodeling Complex to Muscle-Specific Loci. Simone C., Forcales S.V., Hill D., Imbalzano A.L., Latella L., and Puri P.L. Nat Genet 2004 Jul ;36(7):738-43
- Functional interdependence at the chromatin level between the MKK6/p38 and IGF1/Pi3K/AKT pathways during muscle differentiation. Serra C., Palacios D., Mozzetta C Forcales S., Ripani M., Morantte I., Jones D. Du K., Jahla U., Simone C., Puri P.L. Mol Cell 2007 Oct 26 ;28(2):200-13
- TNF/p38 alpha/Polycomb signalling to Pax7 locus in satellite cells links inflammation to the epigenetic control of muscle regeneration. Palacios D., Mozzetta C., Consalvi S., Caretti G., Saccone V., Proserpio V., Marquez V.E., Valente S., Mai A., Forcales S., Sartorelli V., Puri P.L. Cell Stem Cell 2010 Oct 8 ;7(4):455-69
- Signal dependent incorporation of MyoD-BAF60c into Brg1-based SWI/SNF chromatin-remodeling complex. Forcales S., Albini S., Giordani L., Malecova B., Cignolo L., Chernov A., Coutinho P., Saccone V., Consalvi S., Williams R., Wang K., Wu Z., Baranovskaya S., Miller A., Dilworth F., Puri P.L. EMBO J 2012 Jan 18 ;31(2):301-16
4. Epigenetic basis for activation of the myogenic program in ESCs and other pluripotent cell types
Puri lab studied the epigenetic determinants of human embryonic stem cells (hESCs) and induced pluripotent stem cells (hiPSCs) commitment to skeletal myogenesis, by investigating the hESC resistance to direct conversion into skeletal muscle upon ectopic expression of MyoD, which can otherwise reprogram somatic cells into the skeletal muscle lineage. These studies showed that hESC and hiPSC resistance to myogenic conversion is caused by the lack of expression of one structural component of the SWI/SNF chromatin remodelling complex – BAF60C – which is specifically induced in embryoid bodies13. Based on these studies, we have recently established a protocol of hESC-derived 3D contractile myospheres that offers the unprecedented opportunity to dissect and analyze the epigenetic dynamics that underlie the formation of skeletal muscles and to identify changes in the epigenome induced by contractile activity in healthy vs dystrophin-deficient myofibers16,20. We have also determined the identity of the general transcription factors implicated in the activation of skeletal myogenesis17, and we have discovered that replicative senescence is associated with acquisition of resistance to MYOD-mediated activation of muscle gene expression, caused by the constitutive activation of DNA damage repair (DDR) response that impairs cell cycle progression and MYOD activity18. Finally, our recent work has elucidated the mechanism by which MYOD regulates high-order chromatin interactions to define the tri-dimensional (3D) nuclear architecture for the activation of skeletal myogenesis during human somatic cell reprogramming into skeletal muscles19.
- Epigenetic reprogramming of human embryonic stem cells (hESCs) into skeletal muscle cells and generation of contractile myospheres. Albini S., Coutinho P., Malecova B., Giordani L., Savchenko A., Forcales S, Puri P.L. Cell Rep 2013 Mar 28;3(3):661-70
- TBP/TFIID-dependent activation of MyoD target genes in skeletal muscle cells. Malecova B., Dall’Agnese A., Madaro L., Gatto S., Coutinho Toto P., Albini S., Ryan T., Tora L., Puri PL. Elife 2016 Feb 25 ;5:e12534
- 18) DNA damage signaling mediates the functional antagonism between replicative senescence and terminal muscle differentiation. Latella L., Dall’Agnese A, Boscolo F., Nardoni C. Cosentino M., Lahm A, Sacco A., Puri P.L. Genes Dev 2017 Apr 1 ;31(7):648-659
- Transcription Factor-Directed Re-Wiring of Chromatin Architecture for Somatic Cell Nuclear Reprogramming Toward Trans-differentiation. Dall’Agnese A., Caputo L., Nicoletti C., di Iulio J., Schmitt A., Gatto S., Diao Y., Ye Z., Forcato M., Perera R., Bicciato S., Telenti A., Ren B., Puri P.L. Mol Cell 2019 Nov 7 ;76(3):453-472.e8
- Acute conversion of patient-derived Duchenne muscular dystrophy iPSC into myotubes reveals constitutive and inducible over-activation of TGFβ-dependent pro-fibrotic signaling. Caputo L, Granados A, Lenzi J, Rosa A, Ait-Si-Ali A, Puri PL, Albini S. Skelet Muscle 2020 May 2 ;10(1):13
5. Identification, functional, phenotypic and molecular characterization of muscle-interstitial cells – (the fibroadipogenic progenitors – FAPs) in healthy and diseased muscles.
Our work has elucidated the molecular determinants of the interplay between adult muscle stem cells and cellular components of their functional niche (i.e. FAPs), by identifying regulatory networks implicated in compensatory or pathogenic regeneration, and suggesting “disease stage-specific” responses to pharmacological treatment of neuromuscular disorders, such as DMD. Indeed, we have shown that HDACi promote compensatory regeneration and prevent fibro-adipogenic degeneration in mdx mice at early stages of diseases, by targeting a population of muscle interstitial cells – FAPs8 – and have identified a HDAC-regulated network that controls expression of myomiRs and alternative incorporation of BAF60 variants into SWI/SNF complexes to direct the pro-myogenic or fibro-adipogenic FAP activity21. Furthermore, we have recently identified specific subpopulations of FAPs (subFAPs) in physiological conditions and disease22 and we have discovered that specific subFAPs expand and adopt pathogenic phenotypes upon muscle denervation23 or in muscles of patients affected by type 2 diabetes24.
- HDAC-regulated myomiRs control BAF60 variant exchange and direct the functional phenotype of fibro-adipogenic progenitors in dystrophic muscles. Saccone V, Consalvi S, Giordani L, Mozzetta C, Barozzi I, Sandoná M, Ryan T, Rojas-Muñoz A, Madaro L, Fasanaro P, Borsellino G, De Bardi M, Frigè G, Termanini A, Sun X, Rossant J, Bruneau BG, Mercola M, Minucci S, Puri P.L. Genes Dev 2014 Apr 15 ;28(8):841-57
- Spectrum of cellular states within Fibro-Adipogenic Progenitors upon physiological and pathological perturbations of skeletal muscle. Malecova B., Gatto S., Etxaniz U, Passafaro M. Cortez A., Nicoletti C., Giordani L., Torcinaro A., De Bardi M., Bicciato S., De Santa F., Madaro L, Puri PL. Nat Commun 2018 Sep 10 ;9(1):3670
- Denervation-activated STAT3-IL6 signaling in fibro-adipogenic progenitors (FAPs) promotes myofibers atrophy and fibrosis. Madaro L, Passafaro M., Sala D., Etxaniz U., Lugarini F, Proietti D., Alfonsi MV, Nicoletti C., Gatto S., De Bardi M., Rojas-García R, Giordani L., Marinelli S., Pagliarini V., Sette C, Sacco A, Puri PL. Nat Cell Biol 2018 Aug ;20(8):917-927
- Human skeletal muscle CD90+ fibro-adipogenic progenitors are associated with muscle degeneration in type 2 diabetic patients. Farup J, Just J, de Paoli F, Lin L, Jensen JB, Billeskov T, Roman IS, Cömert C, Møller AB, Madaro L, Groppa E, Fred RG, Kampmann U, Gormsen LC, Pedersen SB, Bross P, Stevnsner T, Eldrup N, Pers TH, Rossi FMV, Puri PL, Jessen N. Cell Metab 2021 Nov 2 ;33(11):2201-2214.e11
 Aug 12, 2025
Aug 12, 2025Muscle’s master regulator moonlights as gene silencer
Aug 12, 2025Study uncovers new repressor role for the transcription factor that triggers muscle stem cells to turn into muscle.
 Mar 12, 2025
Mar 12, 2025Meet the La Jolla researcher who helped discover new muscular dystrophy drug
Mar 12, 2025People in Your Neighborhood: Dr. Pier Lorenzo Puri of Sanford Burnham Prebys says he was ‘obsessed with doing something’ to…
 Nov 13, 2024
Nov 13, 2024Decades of dedication led to FDA approval of a new treatment for Duchenne Muscular Dystrophy
Nov 13, 2024Nearly 30 years of discoveries by a Sanford Burnham Prebys scientist and collaborators lead to federal approval of the first…
 Dec 14, 2020
Dec 14, 2020Our top 10 discoveries of 2020
Dec 14, 2020This year required dedication, patience and perseverance as we all adjusted to a new normal—and we’re proud that our scientists
 Sep 14, 2020
Sep 14, 2020Scientists uncover a novel approach to treating Duchenne muscular dystrophy
Sep 14, 2020The study reveals a promising new therapeutic approach for the incurable muscle-wasting condition.
 Sep 10, 2019
Sep 10, 2019How your DNA takes shape makes a big difference in your health
Sep 10, 2019The more we learn about our genome, the more mysteries arise. For example, how can people with the same disease-causing…
Lukas Chavez is an Associate Professor at the Sanford Burnham Prebys. He is also the Director of the Clayes Research Center for Neuro-Oncology at the Institute for Genomic Medicine at the Rady Children’s Hospital, San Diego. In this role, he works with a team of physicians and scientists to capture genomic, transcriptomic, epigenetic and functional data from pediatric brain tumor patients, and uses this information to improve diagnosis and treatment. His research interests focus on structural variants as well as circular extrachromosomal DNA (ecDNA) in childhood cancers. These extrachromosomal DNA circles are frequently found in highly aggressive solid tumors and represent a new target for improved therapeutic approaches.
Education
2010: PhD, Free University, Berlin
Honors and Recognition
2020: St. Baldrick’s Scholar Award, St. Baldrick’s Foundation
2019: Award of Excellence in Pediatric Neuro-Oncology, Society of Neuro-Oncology
2012–2015: Feodor-Lynen Fellowship for Postdoctoral Researchers, Alexander-von-Humboldt Foundation
Related Disease
Brain Cancer, Cancer
Phenomena or Processes
Cancer Biology, Cancer Epigenetics, Chromosome Dynamics, Combinatorial Therapies, Gene Regulation, Genomic Instability, Oncogenes, Transcriptional Regulation
Anatomical Systems and Sites
Brain
Research Models
Clinical and Transitional Research, Computational Modeling, Cultured Cell Lines, Human, Human Cell Lines, Mouse
Techniques and Technologies
Bioinformatics, Cell Biology, Computational Biology, Computational Modeling, Gene Expression, Gene Silencing, Genomics, Single Nucleotide Polymorphisms (SNPs)
 Dec 12, 2025
Dec 12, 2025Sanford Burnham Prebys scientists garner eight cancer research grants from Curebound to advance therapeutic treatments and cures
Dec 12, 2025Ten scientists at Sanford Burnham Prebys Medical Discovery Institute were awarded eight grants yesterday from Curebound, a San Diego-based philanthropic organization
 Aug 15, 2025
Aug 15, 2025Zombie cancer cells give cold shoulder to chemotherapy
Aug 15, 2025Study shows that combining chemotherapy with targeted treatment for these lumbering cells improves outcomes in mice.
 Aug 13, 2025
Aug 13, 2025Chavez joins NIH Cancer Genetics Study Section
Aug 13, 2025Lukas Chavez, PhD, has been named a standing member of the Cancer Genetics Study Section at the National Institutes…
 Apr 14, 2025
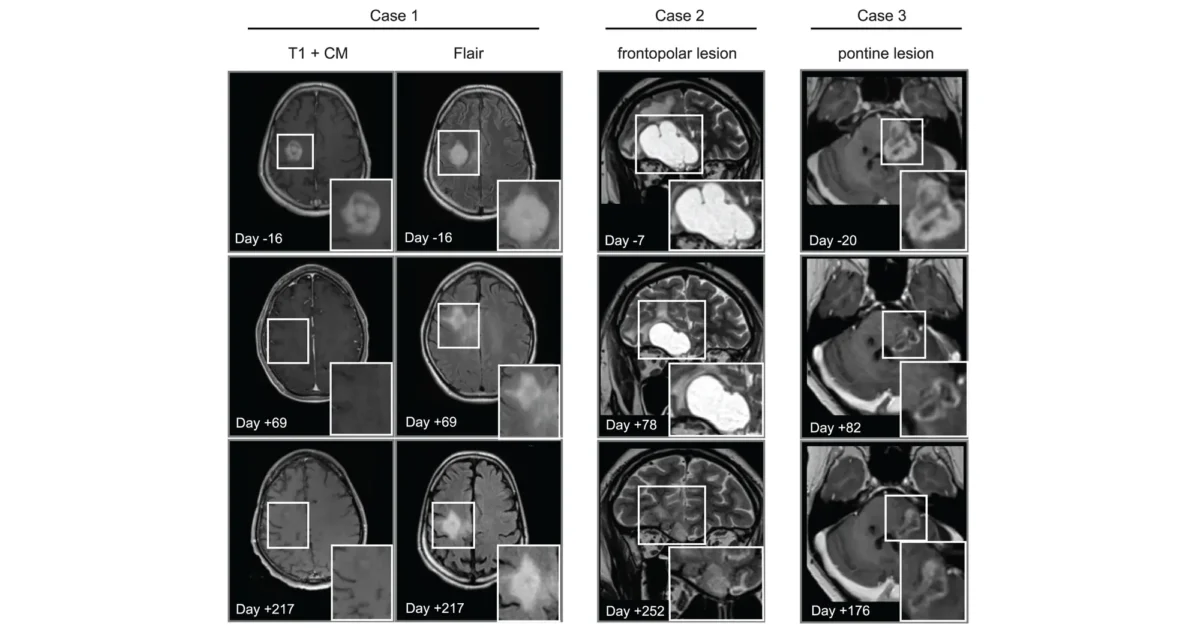
Apr 14, 2025Cancer drug finds new purpose in the brain
Apr 14, 2025Scientists show that a cancer drug travels to and shrinks some brain tumors, which may lead to new therapies.
 Aug 6, 2024
Aug 6, 2024Coding clinic
Aug 6, 2024Rapidly evolving computational tools may unlock vast archives of untapped clinical information—and help solve complex challenges confronting healthcare providers
 Jul 30, 2024
Jul 30, 2024Using machines to personalize patient care
Jul 30, 2024Artificial intelligence (AI) and other computational techniques are aiding scientists and physicians in their quest to create treatments for individuals…