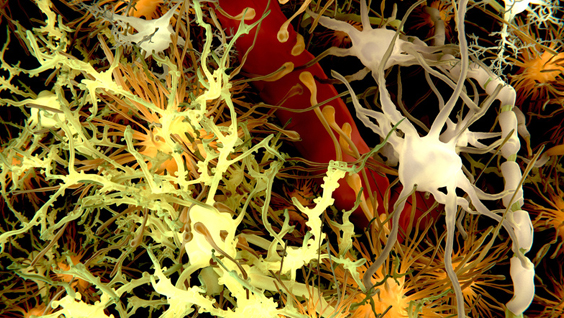
A new study by researchers at Sanford-Burnham reveals the process that leads to changes in the brains of individuals with Down syndrome—the same changes that cause dementia in Alzheimer’s patients. The findings, published in Cell Reports, have important implications for the development of treatments that can prevent damage in neuronal connectivity and brain function in Down syndrome and other neurodevelopmental and neurodegenerative conditions, including Alzheimer’s disease.
Down syndrome is characterized by an extra copy of chromosome 21 and is the most common chromosome abnormality in humans. It occurs in about one per 700 babies in the United States, and is associated with a mild to moderate intellectual disability. Down syndrome is also associated with an increased risk of developing Alzheimer’s disease. By the age of 40, nearly 100 percent of all individuals with Down syndrome develop the changes in the brain associated with Alzheimer’s disease, and approximately 25 percent of people with Down syndrome show signs of Alzheimer’s-type dementia by the age of 35, and 75 percent by age 65. As the life expectancy for people with Down syndrome has increased dramatically in recent years—from 25 in 1983 to 60 today—research aimed to understand the cause of conditions that affect their quality of life are essential.
“Our goal is to understand how the extra copy of chromosome 21 and its genes cause individuals with Down syndrome to have a greatly increased risk of developing dementia,” said Huaxi Xu, PhD, professor in the Degenerative Diseases Program and senior author of the paper. “Our new study reveals how a protein called sorting nexin 27 (SNX27) regulates the generation of beta-amyloid—the main component of the detrimental amyloid plaques found in the brains of people with Down syndrome and Alzheimer’s. The findings are important because they explain how beta-amyloid levels are managed in these individuals.”
Beta-amyloid, plaques, and dementia
Xu’s team found that SNX27 regulates beta-amyloid generation. Beta-amyloid is a sticky protein that’s toxic to neurons. The combination of beta-amyloid and dead neurons form clumps in the brain called plaques. Brain plaques are a pathological hallmark of Alzheimer’s disease and are implicated in the cause of the symptoms of dementia.
“We found that SNX27 reduces beta-amyloid generation through interactions with gamma-secretase—an enzyme that cleaves the beta-amyloid precursor protein to produce beta-amyloid,” said Xin Wang, PhD, a postdoctoral fellow in Xu’s lab and first author of the publication. “When SNX27 interacts with gamma-secretase, the enzyme becomes disabled and cannot produce beta-amyloid. Lower levels of SNX27 lead to increased levels of functional gamma-secretase that in turn lead to increased levels of beta-amyloid.”
SNX27’s role in brain function
Previously, Xu and colleagues found that SNX27-deficient mice shared some characteristics with Down syndrome, and that humans with Down syndrome have significantly lower levels of SNX27. In the brain, SNX27 maintains certain receptors on the cell surface—receptors that are necessary for neurons to fire properly. When levels of SNX27 are reduced, neuron activity is impaired, causing problems with learning and memory. Importantly, the research team found that by adding new copies of the SNX27 gene to the brains of Down syndrome mice, they could repair the memory deficit in the mice.
The researchers went on to reveal how lower levels of SNX27 in Down syndrome are the result of an extra copy of an RNA molecule encoded by chromosome 21 called miRNA-155. miRNA-155 is a small piece of genetic material that doesn’t code for protein, but instead influences the production of SNX27.
With the current study, researchers can piece the entire process together—the extra copy of chromosome 21 causes elevated levels of miRNA-155 that in turn lead to reduced levels of SNX27. Reduced levels of SNX27 lead to an increase in the amount of active gamma-secretase causing an increase in the production of beta-amyloid and the plaques observed in affected individuals.
“We have defined a rather complex mechanism that explains how SNX27 levels indirectly lead to beta-amyloid,” said Xu. “While there may be many factors that contribute to Alzheimer’s characteristics in Down syndrome, our study supports an approach of inhibiting gamma-secretase as a means to prevent the amyloid plaques in the brain found in Down syndrome and Alzheimer’s.”
“Our next step is to develop and implement a screening test to identify molecules that can reduce the levels of miRNA-155 and hence restore the level of SNX27, and find molecules that can enhance the interaction between SNX27 and gamma-secretase. We are working with the Conrad Prebys Center for Chemical Genomics at Sanford-Burnham to achieve this,” added Xu.
# # #
This research was supported in part by US NIH/National Cancer Institute Grant AG038710, AG021173, NS046673, AG030197 and AG044420, and grants from the Alzheimer’s Association, the Global Down Syndrome Foundation, the BrightFocus Foundation (formerly the American Health Assistance Foundation) and the National Natural Science Foundation of China.
To link to the paper click: http://www.cell.com/cell-reports/abstract/S2211-1247(14)00820-1