Xueqin (Sherine) Sun seeks to better understand the genetic and epigenetic underpinnings of cancers, using genome editing technologies, animal and patient-derived models, and other tools to develop more effective cancer therapies.
“My lab is interested in studying how DNA or the machinery that interprets it leads to the transformation of normal cells into cancerous cells and concurrently, their specific vulnerabilities. Identifying these intrinsic vulnerabilities and targeting them properly is profoundly important to developing effective cancer therapies.”
Another aspect of Sun’s work is understanding how cancer cells and tumors change their circumstances and environment to improve survival, including hiding from or repressing the immune system.
“Changes to DNA itself and the way how DNA is interpreted by cells can transform normal cells into cancer cells. And transformed cells propagate by enhancing the misinterpreted DNA information, which in turn becomes the Achilles’ heel of cancer cells. Our goal is to find out how DNA information is misinterpreted in different ways and how to correct it to halt cancer.”
At Sanford Burnham Prebys, Sun and colleagues will employ a host of leading-edge tools and approaches, including functional genomics, artificial intelligence, structural biology, large-scale drug screening, and advanced imaging/spatial technologies.
Sun conducted her postdoctoral fellowship at Cold Spring Harbor Laboratory under the guidance of Alea Mills, PhD, a professor at the National Cancer Institute-designated cancer center at Cold Spring Harbor.
She received her PhD from Wuhan University in China.
Related Disease
Aging-Related Diseases, Brain Cancer, Cancer, Childhood Diseases, Immune Disorders, Inflammatory/Autoimmune Disease, Leukemia/Lymphoma
Phenomena or Processes
Adapter Proteins, Adult/Multipotent Stem Cells, Aging, Angiogenesis, Apoptosis and Cell Death, Bcl-2 Family, Cancer Biology, Cancer Epigenetics, Cell Adhesion and Migration, Cell Biology, Cell Cycle Progression, Cell Differentiation, Cell Motility, Cell Proliferation, Cell Signaling, Cell Surface Receptors, Cellular Senescence, Chromosome Dynamics, Combinatorial Therapies, Cytokines, Development and Differentiation, Disease Therapies, DNA Damage Checkpoint Function, Embryonic/Pluripotent Stem Cells, Epigenetics, Gene Regulation, Genomic Instability, Growth Factors, Hematopoiesis, Host Defense, Host-Pathogen Interactions, Inflammation, Innate Immunity, Kinase Inhibitors, Metastasis, Neurogenesis, Oncogenes, Phosphorylation, Posttranslational Modification, Receptor Tyrosine Kinases, Serine/Threonine Kinases, Signal Transduction, TNF-Family, Transcription Factors, Transcriptional Regulation, Tumor Microenvironment, Tumorigenesis, Tyrosine Kinases, Ubiquitin, Ubiquitin Protease System and Ubiquitin-like Proteins
Anatomical Systems and Sites
Brain, General Cell Biology, Hematopoietic System, Immune System and Inflammation, Nervous System
Research Models
Bacteria, Cultured Cell Lines, Human Adult/Somatic Stem Cells, Human Cell Lines, Mouse, Mouse Cell Lines, Mouse Embryonic Stem Cells, Mouse Somatic Stem Cells, Primary Cells, Primary Human Cells
Techniques and Technologies
3D Image Analysis, 3D Reconstructions, Biochemistry, Bioinformatics, Cell Biology, Cellular and Molecular Imaging, Chemical Biology, Computational Biology, Confocal Microscopy, Correlative Light and Electron Microscopy, Drug Delivery, Drug Discovery, Drug Efficacy, Electron Microscopy, Fluorescence Microscopy, Fragment-Based Drug Design, Gene Expression, Gene Knockout (Complete and Conditional), Gene Silencing, Genetics, Genomics, High Content Imaging, High-Throughput/Robotic Screening, In vivo Modeling, Live Cell Imaging, Live Imaging, Mass Spectrometry, Microscopy and Imaging, Molecular Biology, Molecular Genetics, Nucleic Acid Synthesis, Protein-Protein Interactions, Protein-Small Molecule Interactions, Proteomics, Rational Drug Design, RNA Interference (RNAi), Scanning Cytometry, Small Molecule Compounds, Transgenic Organisms, Transplantation
We seek to understand why cancer occurs and what is the Achille’s heel of cancer, and to develop effective therapeutic interventions.
The successful treatment of any disease requires a good understanding of the mechanisms at work. Cancer is fundamentally caused by aberrant gene expression, which reflects the misinterpretation of DNA information at both genetic and epigenetic levels. We are interested in uncovering DNA-related alterations that drive cancer-favored transcriptional programs, identifying cancer-specific vulnerabilities, and developing effective therapeutic interventions for cancer treatment.
Xueqin Sun’s Research Report
Precise gene expression (the interpretation of DNA) is essential for almost all biological processes, and understanding gene regulation is one of the most pivotal frontiers in biological research under both health and disease circumstances. Gene expression is mainly regulated at genetic (with changes of DNA sequence) and epigenetic (without changing DNA sequence) levels. And gene dysregulation can lead to various health conditions and diseases, including developmental disorders, aging, and cancer. The overarching goal of Sun Lab is to uncover driving genetic and epigenetic alterations involved in cancer, to understand how developmental pathways and aging process impact cancer progression, and to identify mechanisms of action for developing more effective therapeutic strategies.
We are an interdisciplinary lab particularly focused on the following research directions:
- The EP400 chromatin remodeling complex
The EP400 complex is an evolutionarily conserved SWR1-class ATP-dependent chromatin remodeling complex encompassing ~17 components, with a total molecular mass of ~1.5 mega-dalton. The EP400 complex plays critical roles in diverse cellular processes, including chromosome stability, transcription, DNA recombination, DNA damage repair, embryonic stem cell renewal/development, and oncogenesis. The EP400 complex can incorporate histone variants, such as H2AZ and H3.3, into the genome to regulate gene expression. Our recent work discovers BRD8—one of the core subunits of the EP400 complex—as a unique vulnerability of p53 wildtype glioblastoma (GBM), the most prevalent and devastating type of brain cancer. BRD8-driven EP400 complex highjacks H2AZ at p53 target loci to block p53-mediated transactivation and tumor suppression (Nature, 2023). The bromodomain of BRD8 plays the key role in this process. Bromodomain is a druggable domain as evidenced by a number of successful small molecules targeting diverse bromodomains encoded by the human genome across multiple cancer types. Furthermore, findings from others and us suggest that the EP400 complex is involved in different cancers. Thus, we seek to unravel the roles of the EP400 complex in health and disease, and to better understand how to target the EP400 complex for developing effective therapeutic interventions. - The NuRD chromatin remodeling complex
The NuRD complex is also a highly conserved class of ~ 1 MDa multi-subunit chromatin remodeling complexes that consume energy derived from ATP hydrolysis to remodel the configuration of chromatin to control gene transcription programs, with a primary role in gene silencing. Chromatin remodeling is vital for efficiently framing the cellular response to both intrinsic and extrinsic signals and has enormous implications for determining cellular states. NuRD complex is unique in combining ATP-dependent chromatin remodeling, protein deacetylase activity, and recognition of methylated DNA and histone modifications, and has multifarious roles in chromatin organization, transcription regulation, and genome maintenance; thereby, largely impacts health and disease. The NuRD complex has been in the central stage of brain development studies, and is significantly related to brain disorders/diseases. Interestingly, NuRD complex re-assembles by exchanging the chromatin remodeling subunits CHD3/4/5 to achieve specific regulation of an array of genes required for generating distinct cell types in a highly organized manner, especially over brain development. Amongst the genes encoding NuRD complex components, CHD5 is located in human chromosome 1 short arm (1p36), a region that is frequently hemizygously deleted in diverse cancers. Besides genetic deletion, CHD5 is also often silenced in cancer cells due to epigenetic mechanisms, such as promoter hypermethylation, aberrant expression of other chromatin regulators, and microRNAs-mediated translational repression and/or mRNA instability. Our current work seeks to determine whether and how CHD5-driven NuRD complex is involved in tumorigenesis (In preparation, 2024). We will further understand how NuRD complex is involved in both development and tumorigenesis, and identify mechanism of action to develop rational therapeutic strategies. - Novel genetic and epigenetic underpinnings in GBM
GBM is notorious for being a highly complex and plastic cancer type. However, at the genetic level, GBM harbors a relatively low genetic alteration burden compared to the majority of other cancers from pan-cancer profiling studies. This indicates the largely undocumented epigenetic mechanisms that interplay with genetic alterations and co-reprogram transcriptional networks essential for GBM development. Epigenetic changes are usually reversible by nature, as evidenced by numerous successes in targeting epigenetic regulators using small chemical compounds. As actionable therapeutic targets for GBM have been scarce, we are keen to uncover novel epigenetic pathways underlying gliomagenesis under different genetic backgrounds, which will potentially provide promising therapeutic opportunities for GBM treatment. - Novel GBM mouse models
Despite decades of effort, our knowledge about GBM biology is still very limited. GBM harbors a number of genetic alterations. However, among these recurrent genetic lesions, only several have been implicated in gliomagenesis, with most being undocumented. Moreover, the mechanisms by which these genetic alterations are involved in establishing GBM-favored epigenetic landscapes and transcription programs during GBM progression are still largely elusive. The lack of efficient approach to establish mouse models for investigating gene function in gliomagenesis and the limit of current mouse models to recapitulate clinical GBM features in brain is the prime reason that hinders GBM biological research. To this end, we have developed an engineered neural stem cells (NSCs)-based strategy to rapidly generate highly aggressive GBM with desired genetic lesions (genotypes) in mouse brain. Therefore, we will further optimize this strategy to establish a series of novel mouse models possessing recurrent combinations of genetic alterations (genotypes) in GBM, in order to systematically study whether and how these genetic lesions are involved in gliomagenesis and identify genotype-specific dependencies. - Crosstalk between GBM cells and tumor microenvironment
GBM exhibits highly diffuse and infiltrative nature, which contributes to therapeutic resistance and tumor relapse after surgical removal, resulting in dismal prognosis. A better understanding of gliomagenesis involving not only malignant cells themselves, but also the holistic bidirectional interactions of malignant cells with a variety of proximal and distal cells within the organism, is profound for developing novel effective therapies to improve GBM prognosis. Individual invasive GBM cells intermingle with normal brain cells and often cause relapse in brain areas essential for patient survival. Emerging evidence indicates that glioma cells highjack normal brain cells to thrive, and even transform them. However, how gliomagenesis reshapes ecological composition/landscape in host brain and how brain microenvironment affects gliomagenesis are still largely unclear. By using our novel highly invasive mouse models that recapitulate the multiforme diffuse topographies of GBM in brain, we seek to understand the interactions between GBM cells and brain microenvironment, and identify extrinsic pathways that are essential for GBM progression and migration.
Our lab is focused on both fundamental questions in cancer biology and translation of promising therapeutic strategies.
To achieve these, we work together with many fantastic collaborators to develop and leverage cutting-edge technologies, including but not limited to, high-throughput functional genomics (CRISPR/Cas9 screens, exon tiling scan, targeted mutagenesis, etc.), cell and molecular biology, genomics, epigenomics, proteomics, biochemistry, microscopy (2D/3D, time-lapse, two-photon, light sheet, etc.), automated large-scale drug synthesis/screening, structural biology, single cell and spatial multi-omics, artificial intelligence, and bioinformatics. We also establish novel patient-derived models and novel mouse models to facilitate our research programs. Our ultimate goals are to better understand fundamental genetic and epigenetic apparatuses involved in cancer-specific transcriptional networks, provide more effective therapeutic opportunities, and contribute to shifting the paradigms in cancer treatment and precision medicine.
 Aug 19, 2024
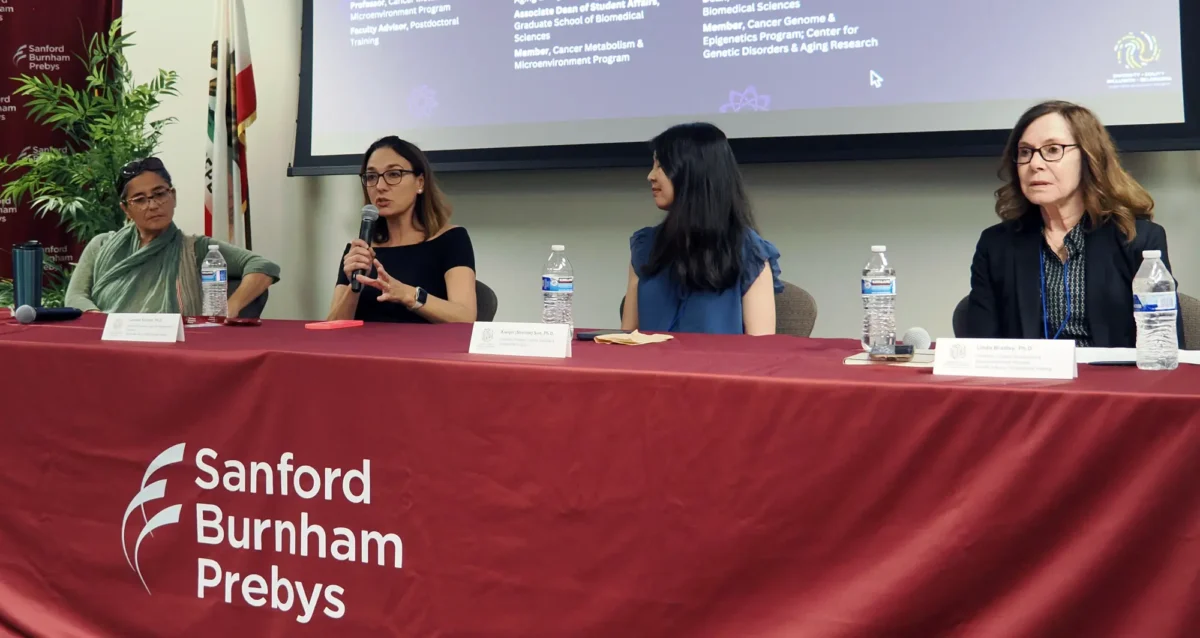
Aug 19, 2024Women in Science event at Sanford Burnham Prebys examines how female faculty members navigate research careers
Aug 19, 2024Topics at the event included work/life balance, caregiving and family obligations, and gender disparities in academic rank at research and…
 Mar 13, 2024
Mar 13, 2024Xueqin Sun seeks to illuminate the underlying causes of cancer
Mar 13, 2024New Sanford Burnham Prebys scientist investigates the mutational powers of cancer cells — and their vulnerabilities
 Dec 19, 2023
Dec 19, 2023Sanford Burnham Prebys continues unprecedented recruitment of early-career scientists
Dec 19, 2023Continuing its rapid and dramatic recruitment of emerging, top-tier researchers, Sanford Burnham Prebys has hired two more highly regarded early-career…