Related Disease
Aging-Related Diseases, Amyotrophic Lateral Sclerosis (Lou Gehrig’s Disease), Arthritis, Cachexia, Inflammatory/Autoimmune Disease, Multiple Sclerosis, Muscular Dystrophy, Myopathy, Neurodegenerative and Neuromuscular Diseases, Sarcopenia/Aging-Related Muscle Atrophy, Spinal Muscular Atrophy
Phenomena or Processes
Adult/Multipotent Stem Cells, Aging, Cell Signaling, Development and Differentiation, Epigenetics, Exercise, Extracellular Matrix, Neurogenesis, Organogenesis, Regenerative Biology, Transcriptional Regulation
Anatomical Systems and Sites
Immune System and Inflammation, Musculoskeletal System, Nervous System
Research Models
Clinical and Transitional Research, Computational Modeling, Human Adult/Somatic Stem Cells, Mouse
Techniques and Technologies
3D Image Analysis, Bioinformatics, Cellular and Molecular Imaging, Gene Knockout (Complete and Conditional), Genomics, High Content Imaging, High-Throughput/Robotic Screening, Live Cell Imaging, Machine Learning, Microscopy and Imaging, Proteomics, Transplantation
The Wang lab is interested in elucidating critical cell-cell interactions that mediate the function of tissue-specific stem cells during regeneration and disease, with a focus on how a coordinated immune response can promote regeneration and how autoimmunity impacts tissue function and hinder repair.
Specifically, the Wang lab aims to identify cellular and molecular crosstalk between muscle, nerve, and immune systems to develop targeted therapies that overcome autoimmune neuromuscular disorders and autoimmune aspects of “inflammaging.”
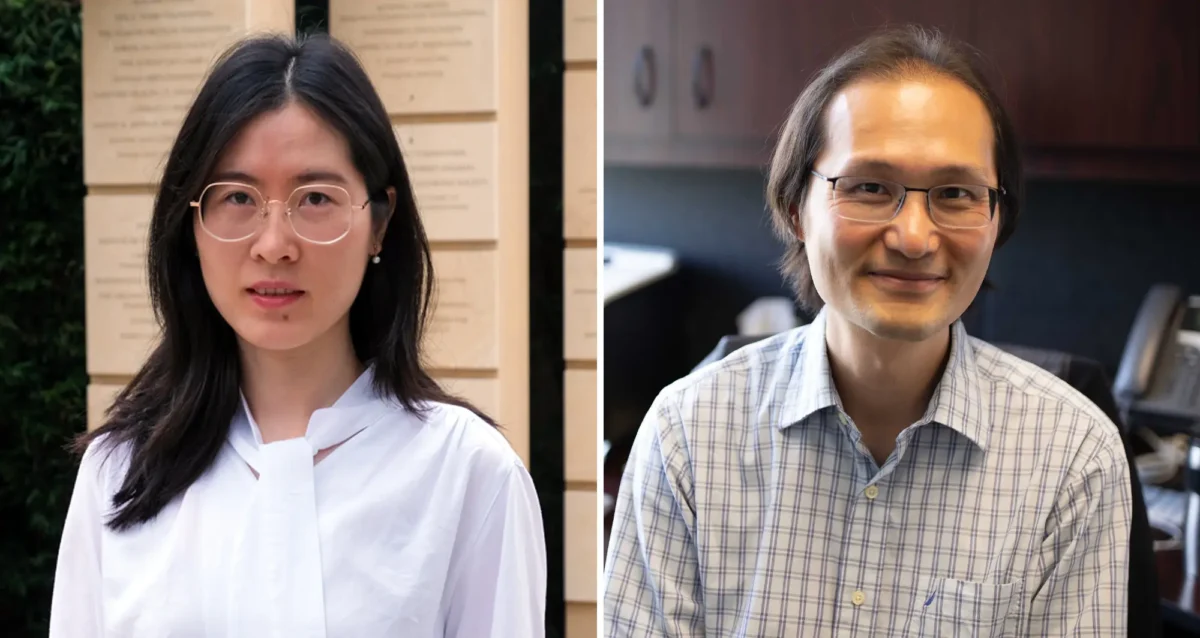
Yu Xin (Will) Wang’s Research Report
The lab’s research is translationally oriented and utilizes interdisciplinary molecular, genetic, computational (machine learning and neural networks), and bioengineering approaches to view biology and disease from new perspectives. We combine multi-omics sequencing and imaging methods to resolve how different cell types work together after injury to repair tissues and restore function. We use a data-driven approach to identify targetable disease mechanisms and, through collaborations with other researchers and clinicians, develop therapies that promote regeneration. Visit our lab website to learn more.